当前位置:
X-MOL 学术
›
J. Chem. Inf. Model.
›
论文详情
Our official English website, www.x-mol.net, welcomes your
feedback! (Note: you will need to create a separate account there.)
MCNN_MC: Computational Prediction of Mitochondrial Carriers and Investigation of Bongkrekic Acid Toxicity Using Protein Language Models and Convolutional Neural Networks
Journal of Chemical Information and Modeling ( IF 5.6 ) Pub Date : 2024-08-12 , DOI: 10.1021/acs.jcim.4c00961 Muhammad Shahid Malik, Yan-Yun Chang, Yu-Chen Liu, Van The Le, Yu-Yen Ou
Journal of Chemical Information and Modeling ( IF 5.6 ) Pub Date : 2024-08-12 , DOI: 10.1021/acs.jcim.4c00961 Muhammad Shahid Malik, Yan-Yun Chang, Yu-Chen Liu, Van The Le, Yu-Yen Ou
|
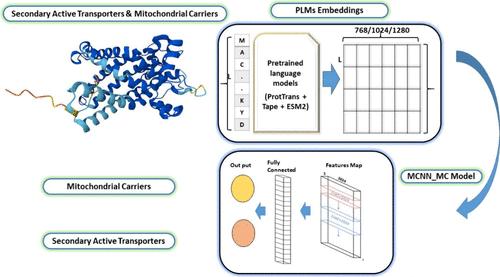
Mitochondrial carriers (MCs) are essential proteins that transport metabolites across mitochondrial membranes and play a critical role in cellular metabolism. ADP/ATP (adenosine diphosphate/adenosine triphosphate) is one of the most important carriers as it contributes to cellular energy production and is susceptible to the powerful toxin bongkrekic acid. This toxin has claimed several lives; for example, a recent foodborne outbreak in Taipei, Taiwan, has caused four deaths and sickened 30 people. The issue of bongkrekic acid poisoning has been a long-standing problem in Indonesia, with reports as early as 1895 detailing numerous deaths from contaminated coconut fermented cakes. In bioinformatics, significant advances have been made in understanding biological processes through computational methods; however, no established computational method has been developed for identifying mitochondrial carriers. We propose a computational bioinformatics approach for predicting MCs from a broader class of secondary active transporters with a focus on the ADP/ATP carrier and its interaction with bongkrekic acid. The proposed model combines protein language models (PLMs) with multiwindow scanning convolutional neural networks (mCNNs). While PLM embeddings capture contextual information within proteins, mCNN scans multiple windows to identify potential binding sites and extract local features. Our results show 96.66% sensitivity, 95.76% specificity, 96.12% accuracy, 91.83% Matthews correlation coefficient (MCC), 94.63% F1-Score, and 98.55% area under the curve (AUC). The results demonstrate the effectiveness of the proposed approach in predicting MCs and elucidating their functions, particularly in the context of bongkrekic acid toxicity. This study presents a valuable approach for identifying novel mitochondrial complexes, characterizing their functional roles, and understanding mitochondrial toxicology mechanisms. Our findings, that utilize computational methods to improve our understanding of cellular processes and drug-target interactions, contribute to the development of therapeutic strategies for mitochondrial disorders, reducing the devastating effects of bongkrekic acid poisoning.
更新日期:2024-08-12































 京公网安备 11010802027423号
京公网安备 11010802027423号