Journal of Molecular Biology ( IF 4.7 ) Pub Date : 2023-03-11 , DOI: 10.1016/j.jmb.2023.168046
Sruthi Sundaresan 1 , Thenmalarchelvi Rathinavelan 1
|
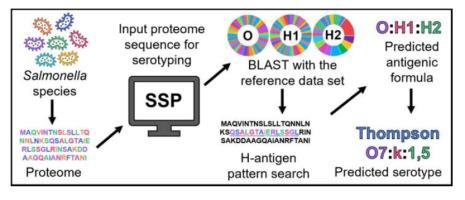
Over 2500 Salmonella species (alternatively, serovars) encompassing different combinations of O-, H1- and H2-antigens are present in nature and cause millions of deaths worldwide every year. Since conventional serotyping is time-consuming, a user-friendly Salmonella species serotyping (SSP) web tool(https://project.iith.ac.in/SSP/) is developed here to predict the serotypes using Salmonella protein(s) or whole proteome sequences. Prior to SSP implementation, a detailed analysis of protein sequences involved in O-antigen biosynthesis and H-antigen formation is carried out to assess their serotype specificity. Intriguingly, the results indicate that WbaP, WecA and GNE initializing transferases can efficiently distinguish the O-antigens, which have Gal, GlcNAc and GalNAc as initial sugars respectively. Rigorous analysis shows that Wzx and Wzy are sufficient to distinguish the O-types. Exceptionally, some situations warrant additional proteins. Thus, 150 additional transferases, RfbE for O2, O9 and O9,46types, Orf17.4 for O3,10 and O1,3,19types, WecB,WbbE and WbbF for O54 and, Wzm and Wzt for O67 are utilized in serotyping. An in-depth analysis of 302 reference datasets representing 56 H1- and 20 H2-types leads to the identification and utilization of 61 unique sequence patterns of FliC and FljB in H-typing. A test dataset of 2136 whole proteome sequences covering 740 Salmonella serovars, including 13 new species and all the O-, H1- and H2-antigens are successfully predicted with 99.72% accuracy. Indeed, SSP also identifies wrongly annotated Salmonella species, hence, can easily identify new species that emerge with any combination of O-, H1- and H2-antigens. Thus, SSP can act as a valuable tool in the surveillance of Salmonella species.
中文翻译:

SSP:使用 O 抗原生物合成蛋白和 H 抗原丝蛋白序列进行沙门氏菌物种血清分型的计算机工具
自然界中存在超过 2500 种包含 O、H1 和 H2 抗原不同组合的沙门氏菌物种(或者血清型),每年在全世界造成数百万人死亡。由于传统的血清分型非常耗时,因此这里开发了一个用户友好的沙门氏菌 物种血清分型(SSP)网络工具(https://project.iith.ac.in/SSP/)来使用沙门氏菌预测血清型蛋白质或整个蛋白质组序列。在实施 SSP 之前,对参与 O 抗原生物合成和 H 抗原形成的蛋白质序列进行详细分析,以评估其血清型特异性。有趣的是,结果表明WbaP、WecA和GNE起始转移酶能够有效地区分分别以Gal、GlcNAc和GalNAc作为起始糖的O-抗原。严格分析表明,Wzx和Wzy足以区分O型。特殊情况下,某些情况需要额外的蛋白质。因此,在血清分型中使用了另外 150 个转移酶,即 O2、O9 和 O9,46 型的 RfbE,O3,10 和 O1,3,19 型的 Orf17.4,O54 的 WecB、WbbE 和 WbbF,以及 O67 的 Wzm 和 Wzt。对代表 56 个 H1 型和 20 个 H2 型的 302 个参考数据集的深入分析导致在 H 分型中识别和利用 FliC 和 FljB 的 61 个独特序列模式。2136 个完整蛋白质组序列的测试数据集,涵盖 740成功预测沙门氏菌血清型,包括 13 个新种以及所有 O-、H1- 和 H2- 抗原,准确率达 99.72%。事实上,SSP 还可以识别错误注释的沙门氏菌物种,因此可以轻松识别 O-、H1- 和 H2- 抗原任意组合出现的新物种。因此,SSP 可以作为沙门氏菌监测的一个有价值的工具。

































 京公网安备 11010802027423号
京公网安备 11010802027423号