Brain Research ( IF 2.7 ) Pub Date : 2021-10-22 , DOI: 10.1016/j.brainres.2021.147702 Jue Liu 1 , Zhifang Deng 1 , Zhijun Yu 2 , Weipin Zhou 2 , Qiong Yuan 2
|
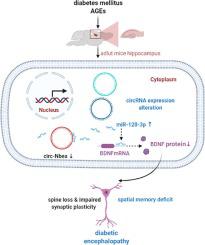
环状 RNA (circRNA) 在人类疾病的各种致病和生物学过程中发挥着关键作用。然而,circRNAs 对糖尿病性脑病 (DE) 发展的影响仍然很大程度上未知。因此,本研究的目的是研究circRNAs表达的变化及其在DE形成中的潜在机制。与db/m 小鼠相比,db/db 小鼠海马区的空间学习/记忆、树突棘和突触可塑性均受损。此外,在用晚期糖基化终产物(AGEs)处理后,神经元的树突棘密度显着降低。我们使用高通量 RNA 测序 (RNA-Seq) 检测 DE 中的 circRNA 表达,结果显示,在 AGEs 处理的原代海马神经元中,183 个 circRNA 发生了显着改变。使用定量实时聚合酶链反应 (qRT-PCR) 选择了三种 circRNA 进行检测,包括 circ-Smox (chr2: 131511984–131516443)、circ-Nbea (mmu-chr3: 56079859–56091120) 和 circ-Setbp1 ( chr18: 79086551–79087180),circ-Nbea 表达显着降低。根据使用 qRT-PCR 和双荧光素酶测定的生物信息学预测和检测,circ-Nbea 海绵 miR-128-3p。基于这些结果,我们推测新发现的 circRNA circ-Nbea 可能在 DE 的发展中发挥重要作用,其机制是通过海绵化 miR-128-3p 介导的。这项研究为 DE 的治疗提供了新的见解。和 circ-Setbp1 (chr18: 79086551–79087180), circ-Nbea 表达显着降低。根据使用 qRT-PCR 和双荧光素酶测定的生物信息学预测和检测,circ-Nbea 海绵 miR-128-3p。基于这些结果,我们推测新发现的 circRNA circ-Nbea 可能在 DE 的发展中发挥重要作用,其机制是通过海绵化 miR-128-3p 介导的。这项研究为 DE 的治疗提供了新的见解。和 circ-Setbp1 (chr18: 79086551–79087180), circ-Nbea 表达显着降低。根据使用 qRT-PCR 和双荧光素酶测定的生物信息学预测和检测,circ-Nbea 海绵 miR-128-3p。基于这些结果,我们推测新发现的 circRNA circ-Nbea 可能在 DE 的发展中发挥重要作用,其机制是通过海绵化 miR-128-3p 介导的。这项研究为 DE 的治疗提供了新的见解。该机制是通过海绵化 miR-128-3p 介导的。这项研究为 DE 的治疗提供了新的见解。该机制是通过海绵化 miR-128-3p 介导的。这项研究为 DE 的治疗提供了新的见解。

"点击查看英文标题和摘要"













































 京公网安备 11010802027423号
京公网安备 11010802027423号